The choice betweenĪ collocated and a staggered formulation is application-dependent. To get a pyvista.UniformGrid which I can then plot with pn.panel (grid, sizingmode stretchboth, height 400, displayslices True) in jupyterlab. I thought pv.DataSetFilters.interpolate would be the right one for this. Them from the collocated to the staggered locations. Hi, I would like to render a volume from a point cloud (polydata). Of the field components by the appropriate phase factors to shift They can also be easily recast on a staggered Yee grid by multiplication The PSATD and PSTD formulations that were just given apply to theįield components located at the nodes of the grid. The electromagnetic fields are solved on a grid, usually using Maxwell’s In the electromagnetic particle-in-cell method (Birdsall and Langdon 1991), smoothing/filtering of the charge/current densities and/or fields on the grid). absorption/emission of particles, addition of external forces to account for accelerator focusing or accelerating component) or numerical effects (e.g. Additional “add-ons” operations are inserted between these core operations to account for additional physics (e.g. The core PIC algorithm involves four operations at each time step: 1) evolve the velocity and position of the particles using the Newton-Lorentz equations, 2) deposit the charge and/or current densities through interpolation from the particles distributions onto the grid, 3) evolve Maxwell’s wave equations (for electromagnetic) or solve Poisson’s equation (for electrostatic) on the grid, 4) interpolate the fields from the grid onto the particles for the next particle push.
9 The Particle-In-Cell (PIC) method follows the evolution of a collection of charged macro-particles (positively charged in blue on the left plot, negatively charged in red) that evolve self-consistently with their electromagnetic (or electrostatic) fields.
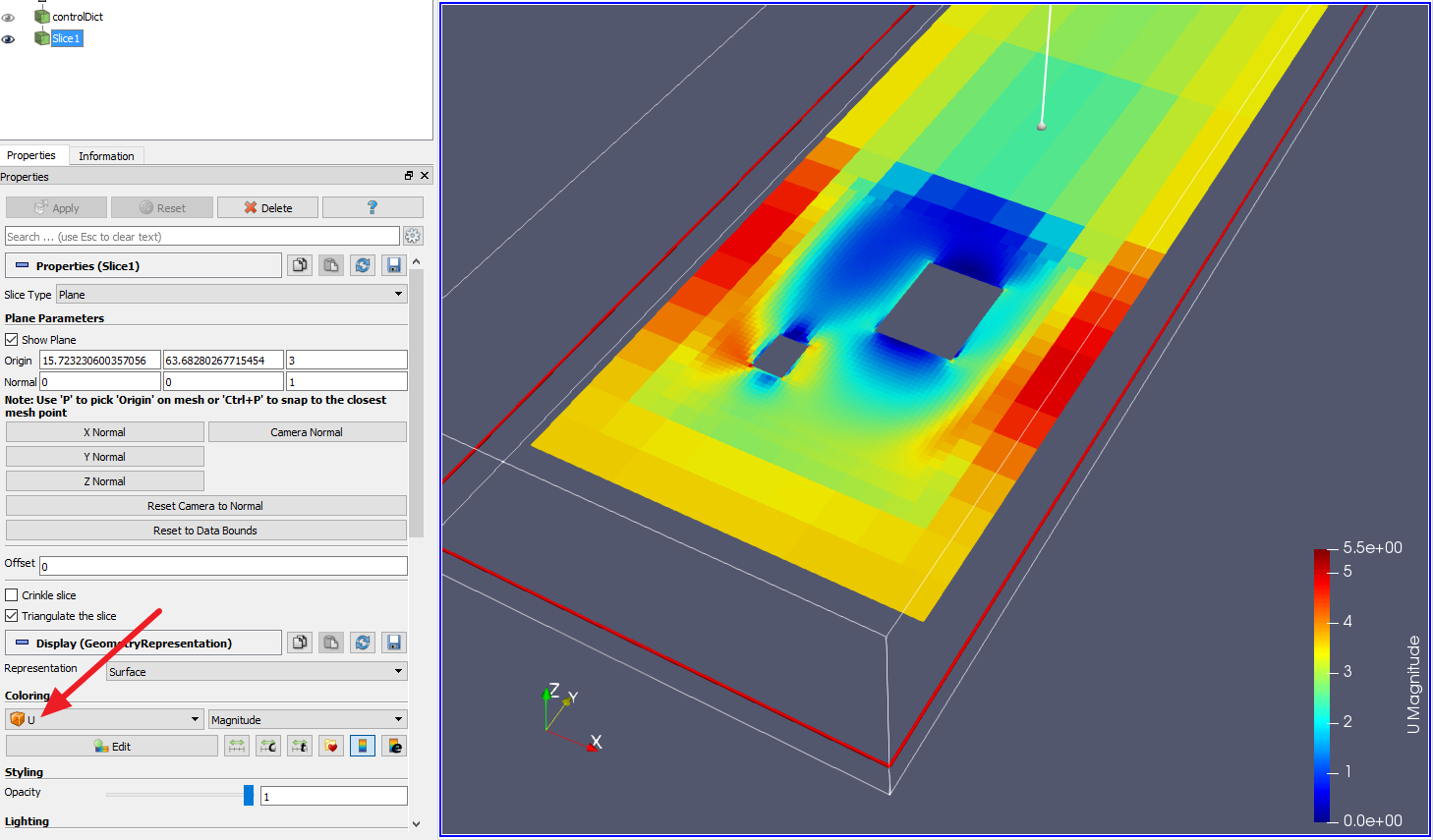
Why go to all this trouble? because doing this will allow you to linearly interpolate your matrices L1 and L2, while still producing an orthonormal rotation. Then, L1=log(O1)=V*log(S)*V' To take the matrix exponential, do the same thing except exp(O1)=V*exp(S)*V1 First, load up the volume and preview it: Note that. For example, let’s slice through the sample geostatistical training image volume. Specifically, do this by taking the eigenvalue decomposition of O, =eig(O), producing a 3x3 matrix V and 3 eigenvalues S. One of the most common slicing filters used in PyVista is the () filter which creates three orthogonal slices through the dataset parallel to the three Cartesian planes. Take the Matrix Logarithm of O1,O2, as L1=log(O1),L2=log(O2). Interpolating the 3x3 rotation is harder, but you can actually pull it off if you have a good matrix library like Eigen. Since the affine matrix has a rotation component (upper left 3x3) and a translation component (right column vector) then to make your new translation component, linearly interpolate the rightmost column. Assuming your matrices are Affine (rotation+translation+scaling) then you can actually do something else.

